The work appears in Nature Communications.
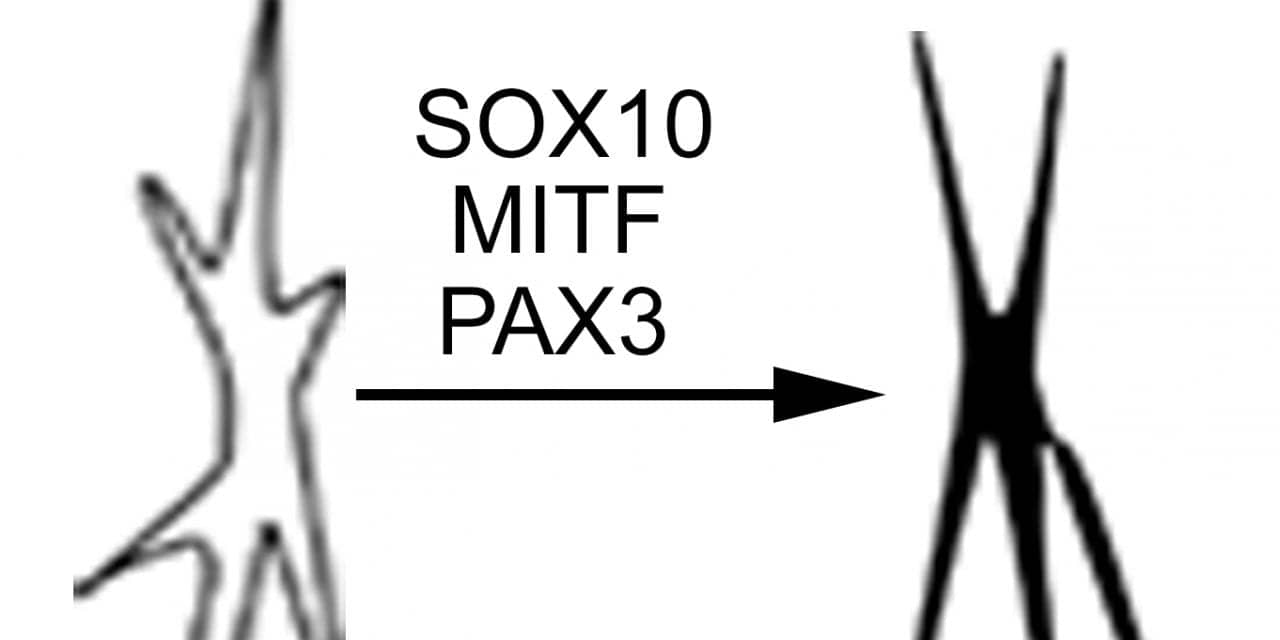
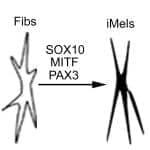
Researchers from the Perelman School of Medicine at the University of Pennsylvania, the Wistar Institute, Boston University School of Medicine, and New Jersey Institute of Technology began by conducting an extensive literature search to identify 10 specific cell transcription factors important for melanocyte development. They then performed a transcription factor screening assay and found three transcription factors out of those 10 that are required for melanocytes: SOX10, MITF, and PAX3, a combination dubbed SMP3.
The researchers first tested the SMP3 combination in mouse embryonic fibroblasts, which then quickly displayed melanocytic markers. Their next step used a human-derived SMP3 combination in human fetal dermal cells, and again melanocytes (human-induced melanocytes, or hiMels) rapidly appeared. Further testing confirmed that these hiMels indeed functioned as normal melanocytes, not only in cell culture but also in animals—using a hair-patch assay—in which the hiMels generated melanin pigment. The hiMels proved to be functionally identical in every respect to normal melanocytes, the study showed.
Xu and his colleagues anticipate using their new technique in the treatment of a wide variety of skin diseases, particularly those such as vitiligo, for which cell-based therapies are the best and most efficient approach.
The method could also provide a new way to study melanoma. By generating melanocytes from the fibroblasts of melanoma patients, “we can screen not only to find why these patients easily develop melanoma, but possibly use their cells to screen for small compounds that can prevent melanoma from happening,” said Xiaowei “George” Xu, MD, PhD, an associate professor of pathology and laboratory medicine at Perelman School of Medicine at the University of Pennsylvania, in a news release.


